The kyphosis data in the data file
has already been used in the Bayesian model selection example in the MCMC notes. Here we do a frequentist analysis. The R statements below do a logistic regression (with the default logit link function).
The regression coefficient for the predictor Age,
which is 0.010929 with a reported standard error of 0.006416,
does not appear to be statistically significant (P = 0.08849).
But that standard error is derived from Fisher information and relies
on the validity of the usual asymptotics of maximum likelihood
.
What if n isn't large enough? What should we really think?
Here's a parametric bootstrap calculation of this P-value.
p.hat <- predict(out.little, type = "response")
If we left off the type = "response" we would get linear
predictor values rather than expected values, which is not what is wanted.
The simulation is quite trivial
present.star <- as.numeric(runif(n) < p.hat)
Another way to do the same thing would be
present.star <- sample(0:1, prob = p.hat, replace = TRUE)
Other probability models are harder, but the principle is the same. Simulate the null model with estimated values of the parameters (in the null model) plugged in.
Because the Monte Carlo error was too big to tell any difference between the parametric bootstrap P-value and the asymptotic P-value, we do another bootstrap with a bigger bootstrap sample size.
Since it takes way too long to do in Rweb, the results are in the file
(the input file was parm.R).
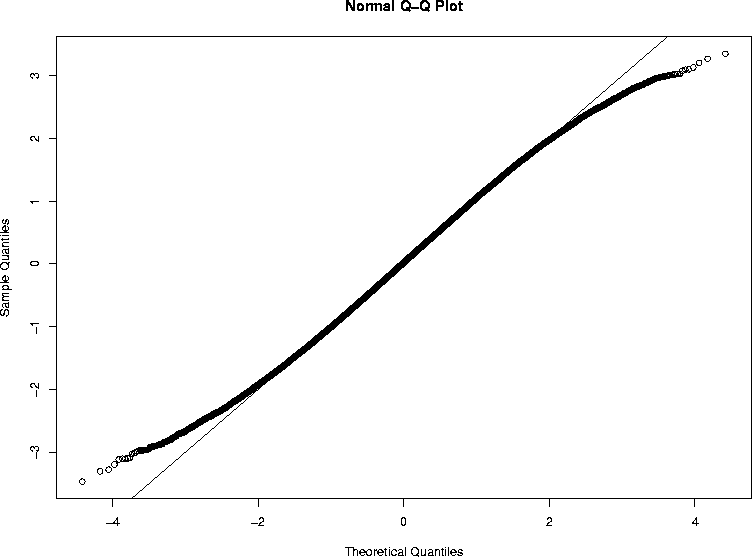
The plot made by this run (and turned into GIF format by the same magic that Rweb uses) is shown below.

glm
function starts generating warnings about nonconvergence. This is
not really the fault of the programmers of the glm function.
The whole GLM community has always been extremely lackadaisical about
convergence issues and existence of maximum likelihood estimates.
It is a flaw of the glm function that it does not return
an error code. Just printing a warning but doing nothing that the
users code can respond to is extremely frustrating. Programmer hostile!
While I was ranting about this, Murali Haran told me about the
try function, that does allow one to respond to errors.
So turn the warnings into errors. That's what
options(warn = 2)
does. And then catch the errors with the try function.
That's what
out.star <- try(glm(present.star ~ Age + Number + Start, family=binomial))
if (! inherits(out.star, "try-error")) {
coef.star <- summary(out.star)$coefficients
z.star[i] <- coef.star["Age", "z value"]
}
does.
The try function just does nothing if there is no error
(its result is the result of its argument). If there is an error, then
the result will inherit
from class "try-error"
(and any result of the argument of the try function is lost)
Thus we can do what we would ordinarily do if
inherits(out.star, "try-error") is FALSE.
Otherwise, we do nothing and z.star[i] is unchanged from
its initial value, which is NA.
NA values.
Since these reflect programmer brain damage (and ultimately theoretician
brain damage), we can hardly count NA values as evidence in
favor of the hypothesis we favor. Thus we should count them in favor of
the null hypothesis for a usual test and in favor of the alternative for
a goodness of fit test. Here we count them in favor of the null,
that is, we include them in the numerator of the bootstrap P-value
p.val <- (sum(abs(z.star) >= z.hat, na.rm = TRUE) +
sum(is.na(z.star)) + 1) / (nboot + 1)
They go bad right around theoretical quantile
plus or minus 2,
which is the 0.025 level one-tailed or the 0.05 level two-tailed.
So we would need the parametric bootstrap for this calculation if the
results were more statistically significant!